Impact
How the Julich Brain Atlas contributes to advancing neuroscience, medicine and technology
The Julich Brain Atlas has become a modern-day reference of the brain for the neuroimaging community. Its impact also includes brain medicine. For example, clinical researchers have been using the high-precision data of the BigBrain to optimise surgery for the treatment of epilepsy. It has also contributed to advances in supercomputing and AI not least because the analysis of entire brains at cellular resolution produces massive amounts of data and poses new challenges to high-performance computing.
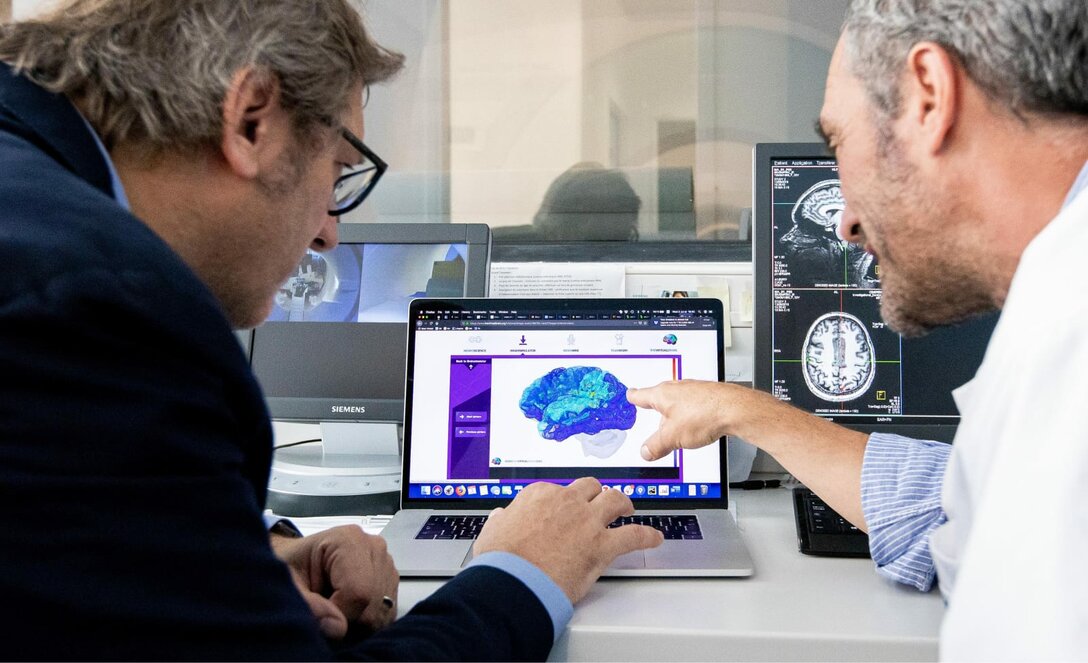
Improving epilepsy surgery with digital brain models
In millions of epilepsy patients, pharmacological treatment is not effective and surgical intervention is the only treatment option. Within the Human Brain Project (HBP), scientists from France have developed personalised brain models (‘The Virtual Brain’) to identify the areas where seizures emerge in a patient’s brain. With the help of the Julich Brain Atlas the accuracy of the brain models is being improved. A 400-patient clinical trial is currently ongoing with the aim of providing surgeons with a precise tool to help individual surgery decisions and improve outcomes.
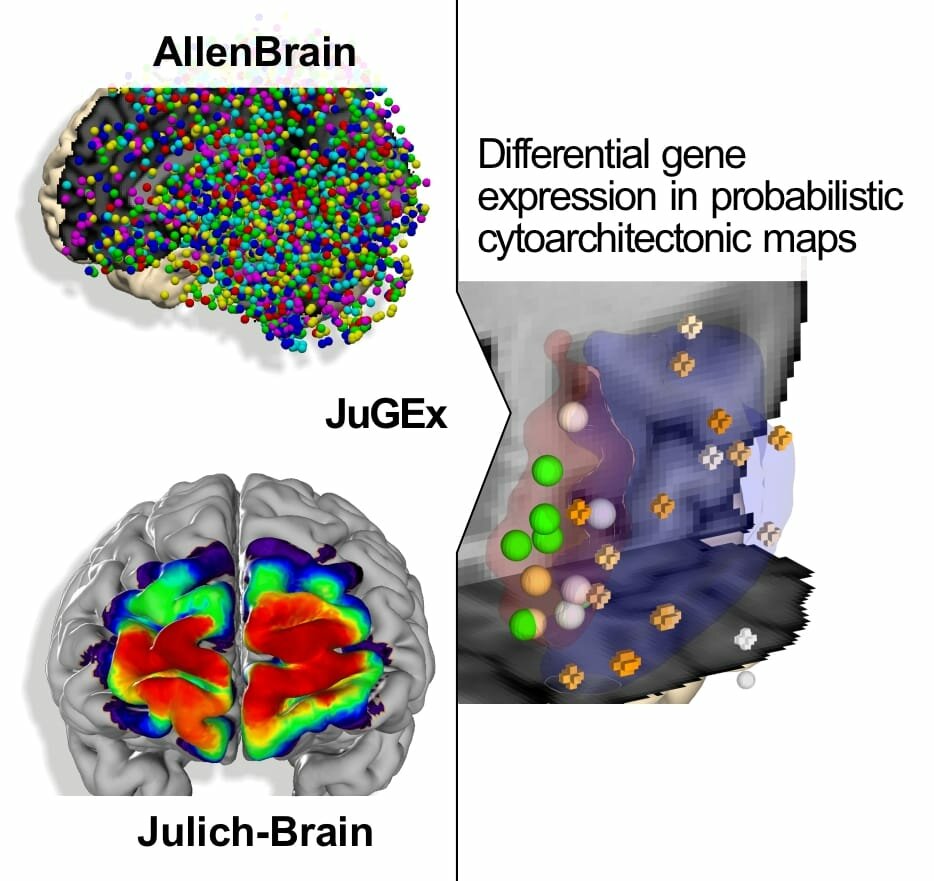
© Bludau et al. Brain Struct Funct 223, 2335–2342 (2018). https://doi.org/10.1007/s00429-018-1620-6
Linking cytoarchitecture to gene expression
We have developed the JuGEx tool to allow integrating information on brain architecture from the Julich Brain Atlas with gene expression data from the Allen Human Brain Atlas. The tool enables detailed insights into how areas with specific gene activities and microanatomical architectures contribute to brain function and dysfunction. Using JuGEx, we found differences in the expression of several candidate genes for major depressive disorder in a disease-affected part of the brain. In this way, the Julich Brain Atlas helps researchers to better understand disease.
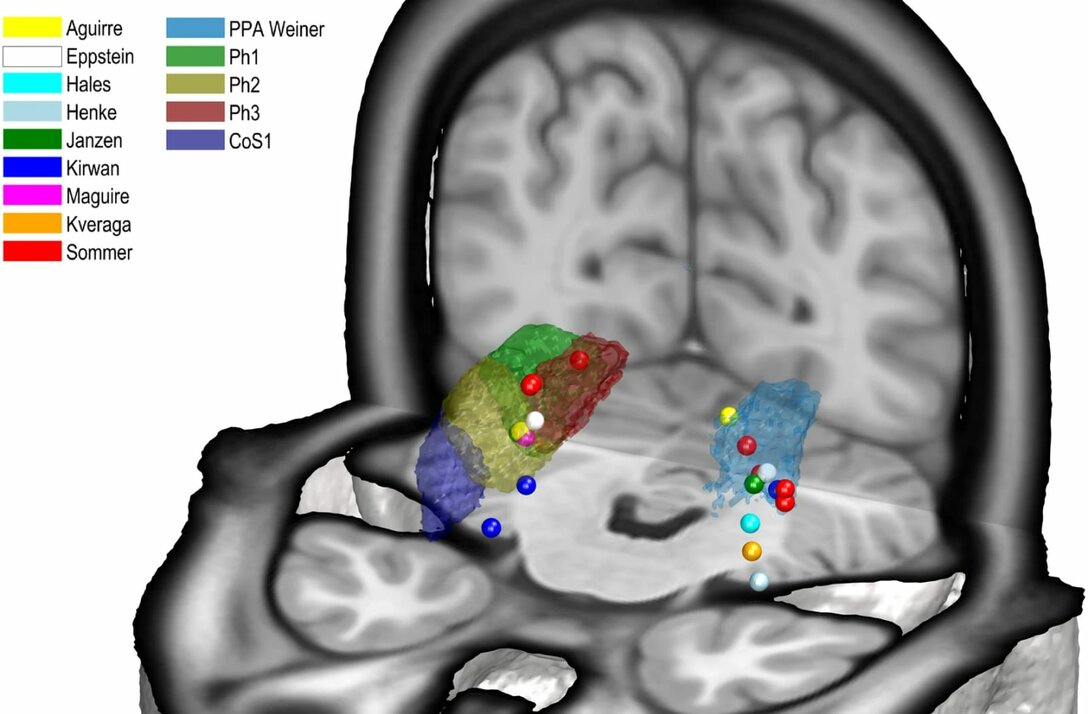
Correlating microstructure and function
The Julich Brain Atlas enables researchers to directly relate microstructural segregation of brain areas to functional imaging data. A recent example is a study in which we have identified four new areas in the parahippocampal cortex and were able to demonstrate that their distinctions in cytoarchitecture correlate with different functions in visual-spatial orientation and associative memory. This finding is based on superimposing probability maps of the new areas with activation peaks from in vivo neuroimaging (see picture).
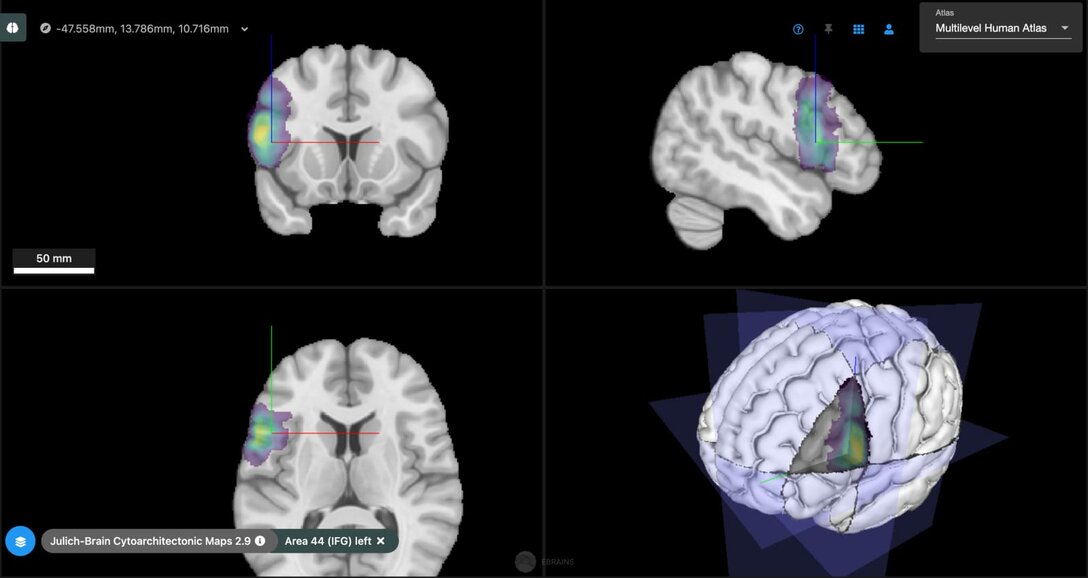
Explaining individual differences during aging
The Julich Brain Atlas also helps to explain variability of cognitive, lifestyle and neurodegeneration profiles between different individuals. We have recently shown how such phenotype data can be integrated with regional genetic, molecular and connectional data using the Julich Brain Atlas via the HBP’s EBRAINS infrastructure to explain individual differences in neurodegeneration.